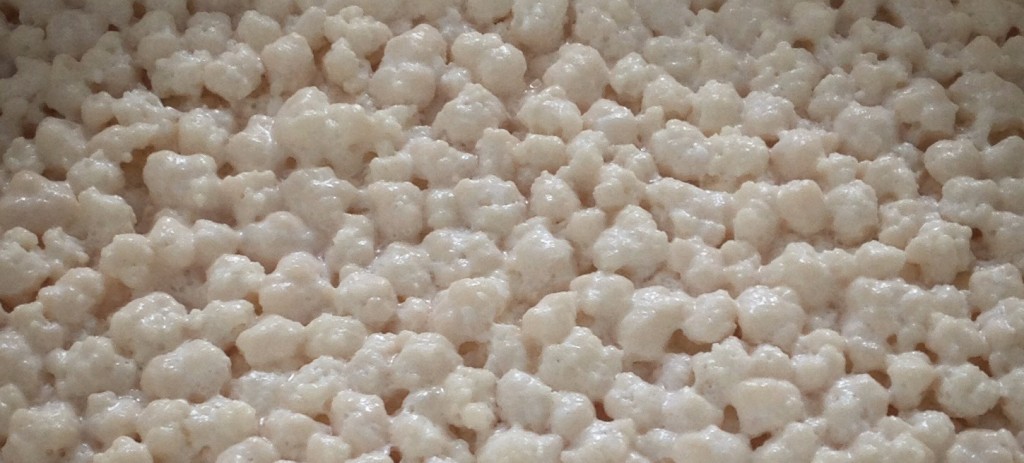

Dissecting the Microbial Diversity of Kefir
Kefir is a thick, sour, and sometimes slightly spritzy fermented milk drink produced through the action of the bacteria and fungi within kefir ‘grains’, a classic example of a SCOBY (Symbiotic Community of Bacteria and Yeasts). Despite a history that dates back several millennia, kefir and the microbes that produce it remain little-understood. Two recent papers from China and Ireland set out to explore the microbial diversity of kefir samples collected from a wide geographical area. One also provides insight into the physical structure of the kefir grain, and the distribution of yeast and bacteria across it.
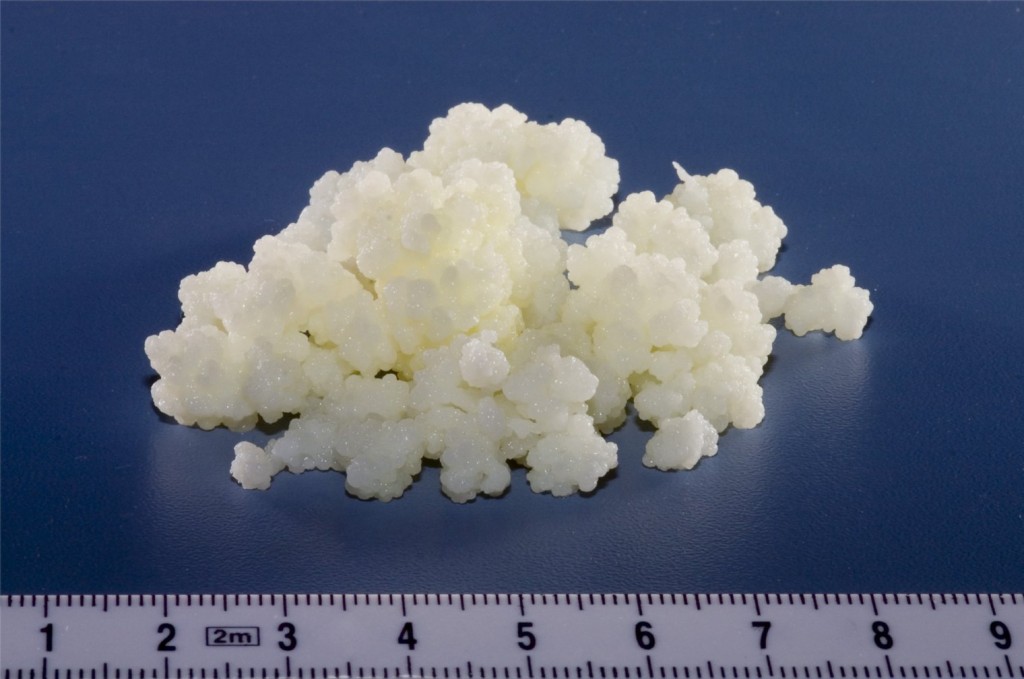

Grains of kefir, a fermented dairy beverage. Photo from Wikipedia user A. Kniesel.
Research Questions That Were Asked in These Studies
What can microscopy tell us about the physical structure of kefir grains and the organisms that live on and within them? What are the prominent fungal genera within three Tibetan kefir cultures, and how stable are these populations over time?
What can high-throughput molecular methods tell us about the bacterial and fungal diversity of kefir cultures from around Western Europe? Within each European kefir sample, how does the microbial composition of the kefir grain compare to the cultured milk it produces?
Methods Used by the Researchers
The team that analysed the grains’ structure used microscopy at a range of resolutions, and various types of fluorescent staining, both general and specific to individual species, to identify and locate specific microbes within their samples.
Both groups used molecular methods for identifying yeasts (and in the case of the Irish team, bacteria) within their samples. The primary advantage of these molecular methods (as opposed to traditional culture-based methods) is that they don’t rely on whether the microbes will happily grow on laboratory media. With many thousands of reads from a single sample, the ‘high-throughput’ methods used by the Irish team gave a finely-detailed view of what was in it, and in what quantity.
Summary of Findings
Lu et al. found that the cauliflower-like kefir grains are formed of many globular, hollow ‘small grain units’ which may be shed from the larger community and regenerate new colonies. The the walls of these hollow grains are made up of rod-shaped bacteria, which form a ‘polyhedron-like net structure’ into which yeasts were embedded, primarily on the outer surface of the globules. Marsh et al. cite studies suggesting that within a 24-hour fermented kefir culture, bacteria outnumber yeasts by a factor of around 100 to 1.

Scanning Electron Micrograph of Tibetan kefir grains from Lu et al. (2014). A+B: outer surface of grains; C+D: cross-section of grains; E-H inner surfaces of grains cultured normally and aseptically. See full information in original paper (link below).
Fungal Communities: The two groups reported different fungal populations in the European versus Tibetan kefir grains. Both studies found that the overwhelming majority of fungi belonged to a family of budding yeasts called Saccharomycetaceae, but while the Chinese group reported a dominance of Saccharomyces cerevisiae across their samples, the Irish group found it only in minute quantities in one of their 25 European kefir cultures. Likewise, the Chinese group failed to find yeasts of the genus Kazachstania in their cultures, whereas the Irish group reported that it was the most common fungal genus across all their samples. However, it is important to note that a large part of this reported variation is likely to be down to differences in classification and nomenclature. Indeed, Marsh et al. note that the relationship between Kazachstania and another frequently-identified yeast, Naumovozyma, is extremely close, and in the not-too-distant past they both would have been considered part of the genus Saccharomyces.
Bacterial Communities: Only the Irish group looked into the bacterial makeup of their kefir cultures. They found that the cultures were dominated by Firmicutes (mainly lactic acid bacteria, including Lactobacillus) and Proteobacteria (mainly Acetobacter). The dominant genus in the kefir grains was Lactobacillus, followed by Acetobacter. In the kefir milks produced by that same set of grains, however, the diversity of bacteria was lower than the grains, and the lactic acid bacterium Lactococcus dominated.The proportions of different bacteria varied considerably amongst the individual cultures.
Microbial Diversity and Geographical Distribution: Neither group found that the kefir cultures were very diverse. On a species level, the Irish group found relatively low alpha-diversity (diversity within a given sample) of bacteria and fungi. Interestingly, the bacterial diversity of the grains was somewhat greater than the cultured milk that they produced, but the fungal diversity of the cultured milk was higher than the commensurate grains. Counterintuitively, statistical analysis showed that there was also no significant correlation between the milks and their grains: ‘the similarities between kefir grain communities were not the same as the similarities between kefir [milk] communities.’
The Irish group did not find a significant correlation between geography and microbial composition of the kefir communities that they studied.
Practical Implications
Despite a perhaps-disappointing lack of clear geographical distribution patterns, low species diversity of kefir cultures, or even correlations between the dominant communities in grains and cultured milks, these two papers do point out some interesting phenomena with practical ramifications for the kefir enthusiast:
Different organisms prosper and dominate in different environments: the milks and grains often contained organisms that were not identified in the corresponding mother culture or fermentation product, even using high-throughput (highly-sensitive) methods. This reflects an overarching trend in microbial foods of all kinds: infinitessimal amounts of an organism can grow to significant levels in a short period of time, and sometimes it only takes a small change in an environment to shift the patterns of growth.
The dominance of one or another genus in one culture versus another seems to be closely linked with the physiochemical and nutritional conditions in which the culture has developed over time. Medium is important: the Chinese group found that one organism, Yarrowia lipolytica, grew from ‘nearly zero to being the dominant population’ in one of their three samples over the course of 10 months of culture in the lab. The scientists hypothesized that this shift was due to their choice to culture it in whole milk rather than the skimmed milk medium used for its original propagation; because Y. lipolytica can metabolise fats but not lactose, the whole milk culture gave it the nutrients it needed to prosper and eventually dominate.
The lack of significant geographical differences between kefir cultures in terms of yeast or bacteria follows the same trend identified by Wolfe et al. in their survey of cheese rinds. Could this be a widespread microbial foods trend?
The Irish group was interested in exploring kefir as a ‘potential delivery system for viable health-promoting organisms to the gut.’ Bifidobacteria have received a lot of attention recently as a potentially important probiotic, but it was clear from the group’s findings that Bifidobacteria are rarely found at detectable levels in kefir, and that they do not compete well in a kefir environment. However, apart from any potential direct probiotic effect, the organic acids and bioactive peptides produced during the fermentation may also promote health benefits—and moreover, hopefully, the kefir produced is also delicious!
Both papers referred to the fact that simply mixing and culturing the species associated with kefir grains together doesn’t result in the formation of a stable kefir SCOBY; kefir likely evolved ‘from a long-term symbiotic interaction’ between the yeasts and bacteria. There is clearly still much to be discovered about these elegantly co-evolved microbial communities.
Post written by Bronwen Percival
For more details on these studies, please check out the full articles here:
Lu et al. (2014) Fine Structure of Tibetan Kefir Grains and Their Yeast Distribution, Diversity, and Shift: PloS ONE 9(6): e101387. Full text here.
Marsh et al. (2013) Sequencing-Based Analysis of the Bacterial and Fungal Composition of Kefir Grains and Milks from Multiple Sources. PloS ONE 8(7): e69371. Full text here.
Very Nice write-up! Thank you.
[…] need to buy a microscope, others have done it for you: Science Digested: Dissecting the Microbial Diversity of Kefir – MicrobialFoods.org For coconut milk kefir, I use organic canned coconut milk, of the sort you'd cook with. Thai […]
I think there is a spelling mistake here: “because Y. lipolytica can metabolise fats but not lactose, the whole milk culture gave it the nutrients it needed to prosper and eventually dominate.” Did you mean to write “gave in”? Because it’s not clear in this way. Thank you very much.
No the sentence sounds correct. The whole milk have IT the fat it needed to prosper that the skim milk couldn’t.
Am very interested in growing kefir grains to share with friends,but my kefir while it ferments , the grains have remained the same quantity. Help!
Hi Saro –
I am not exactly sure why you aren’t getting growth with your kefir. How long have you had the grains? What type of milk are you using? What temperature?
Best,
Ben
You need to use the same growing medium as the one the grains were grown in. A coconut milk kefir may not grow well in cow milk, and vice versa.
Does this article imply that my kefir does not have the amount of beneficial microbes that are talked about. Don
Being an RN in a nursing home we have residents with chronic wounds. One Dr agreed to us using Kefir which we applied daily for a couple of weeks. This cleared the strong mixed bacteria field of the ulcer can gave us a clean base to then dress. After many years of traditional antibiotics unsuccessfully used the clear the bacterial film on this ulcer kefir worked. The residents ulcers on both legs are now healed! Just amazing really.
That is an amazing point about kefir, I did not know!
Deborah, the wounds you are speaking of, were they staph infections?
Hi,
How much temperature and time do you recommed to use for drinks made by kefir (pasteuriation), i see there are a lot of microorganism in it. what are the most resistant ???
Thanks!!
If you pasteurize it you are eliminating the healthy bacterial and yeast benefits of the kefir. The point of consuming a live culture is that the bacteria is beneficial to your gut.
The studies indicate that the Bifidobacterium genus does not compete well in a water kefir. Are there any fermented drinks you know of that have Bifidobacterium as a sizeable percentage of the ferment?
This is an interesting question. It seems like most fermented foods have very small amounts of bifidobacteria, if any… since pre-industrial humans likely had plenty of Bifidobacteria in their guts, this probably didn’t come up in traditional situations.
I’ll look into it more, but wonder if you’d have to artificially increase the Bifidobacteria if you want it to be a big part. This is done with yogurt. You could probably also do it with sour poi if you want something traditionally dairy free.
I also wonder if Bifidobacteria would become dominant in a water kefir culture that includes oligosaccharides, which is their preferred diet. Maybe you could add a dose of inulin FOS powder or something like that? It seems clear from the study that the Bifs are likely there – just not evident.
Hi,
I am wondering if you can advise me. I am making kefir from culture and am under the impression that if I let it ferment long enough, grains will begin to grow, is that correct?
thanks
Lindsey